St. Jude Family of Websites
Explore our cutting edge research, world-class patient care, career opportunities and more.
St. Jude Children's Research Hospital Home

- Fundraising
St. Jude Family of Websites
Explore our cutting edge research, world-class patient care, career opportunities and more.
St. Jude Children's Research Hospital Home

- Fundraising
Loeffler Lab
Deciphering the mechanisms of self-renewal and fate decisions in normal and mutant hematopoietic stem cells using quantitative single-cell dynamics
About the Loeffler Lab
As stem cells in the bone marrow divide and age, they progressively lose their ability to self-renew. Our laboratory is interested in understanding what mechanisms control this phenomenon. We combine novel quantitative single-cell live-cell imaging, microfluidics, robotics, and cutting-edge sequencing approaches to decipher the epigenetic, genetic, and molecular dynamics of stem cell fate regulation. Ultimately, this work will enhance our fundamental understanding of the hematopoietic system in development, and dysfunction.
Our research summary
Hematopoietic stem cells (HSCs) can self-renew and differentiate to regenerate the hematopoietic system. While this regenerative potential is extensive, HSC function declines with successive rounds of divisions and increasing age. The molecular mechanisms for this decline remain mysterious and poorly understood. Our goal is to better define the molecular mechanisms that govern stemness and differentiation in the hematopoietic system and how these processes are altered in early stages of hematological diseases.
Clonal Hematopoiesis
To understand what drives this loss of self-renewal potential, our laboratory uses clonal hematopoiesis as a model. Stem cells with mutations in genes associated with clonal hematopoiesis acquire an advantage in fitness relative to wildtype HSCs. Mutant HSC clones expand over time, and eventually acquire additional mutations and develop into severe hematological and non-hematological diseases. This process often takes decades and mutant clones remain undetected, because the early molecular changes in mutant HSCs are subtle with minor alterations in cell behavior and function. It has thus been difficult to decipher the underlying molecular mechanisms of clonal hematopoiesis using classical molecular biological tools, which are often semi-quantitative and rely on population averages or single-cell analysis at discrete time points. Evidence suggests that epigenetic changes contribute to the early clonal expansion, but the underlying molecular details remain unclear and require further investigation using novel quantitative tools with single-cell resolution and sufficient sensitivity to detect subtle changes even at the very early stages of disease.
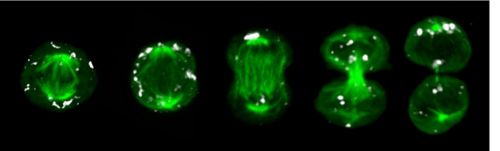
Quantification of dynamic molecular processes at the single-cell level
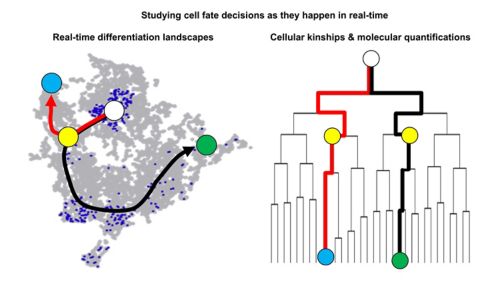
Leveraging continuous quantitative time lapse imagine, microfluidics, robotics, next generation sequencing and bioinformatics we can monitor and manipulate normal and mutant stem cell behavior in extended time course experiments at single-cell resolution. We can track individual HSCs and their offspring over many cell generations and quantify changes in cell morphology, migration, adhesion, transcription factor levels, signaling pathway activities, epigenetics and inheritance during cell division using dynamic reporter constructs. This approach enables us to observe cell fate decisions in real-time as they happen and facilitates to study cellular responses in changing environments to establish causality.
Our work is interdisciplinary and at the interface between stem cell biology, bioinformatics, and cellular engineering. We develop and combine novel cutting-edge bioimaging with next generation multi-omics to ask fundamental questions about stem cell biology, the earliest steps of disease development and the molecular reasons of aging. The answers to these questions have the potential to shift paradigms and lead to novel, improved therapies. We are actively seeking highly motivated scientists and trainees with expertise in molecular biology, cell biology, genomics, bioinformatics or related fields. Candidates with an interest in stem cell biology, hematological malignancies and aging that are motivated to work in an interdisciplinary environment, curious and eager to integrate single-cell analysis, live-cell imaging, bioinformatics, and engineering into their project portfolios are encouraged to apply.
Selected Publications
Contact us
Dirk Loffler, PhD
Assistant Member, Hematology
MS 341
St. Jude Children's Research Hospital

Memphis, TN, 38105-3678 USA GET DIRECTIONS