St. Jude Family of Websites
Explore our cutting edge research, world-class patient care, career opportunities and more.
St. Jude Children's Research Hospital Home

- Fundraising
St. Jude Family of Websites
Explore our cutting edge research, world-class patient care, career opportunities and more.
St. Jude Children's Research Hospital Home

- Fundraising
Center for Spatial Omics
Spatial omics is revolutionizing biomedical research by uniting genomics, proteomics, and transcriptomics to define cells within the context of their spatial environment.
OVERVIEW
The Center for Spatial Omics (CSO) at St. Jude, established as part of the 2022-2027 Strategic Plan, is a shared resource that uses advanced spatial omics platforms to study cell interactions and organization in tissues at single-cell resolution while maintaining tissue architecture. The CSO utilizes various spatial omics assays and instrumentation to advance discoveries in catastrophic childhood diseases.
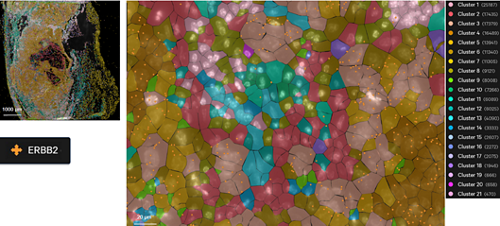

Fluorescence signal (left) and assignment of individual transcripts by cell segmentation (right) via RNA/ISH-based Xenium methodology performed on human breast cancer sample.
IMPACT
The Center of Spatial Omics (CSO) enables high-resolution, highly multiplexed spatial analysis of gene and protein expression within intact tissues, providing St. Jude investigators with access to molecular information at a scale and level of context not achievable with conventional approaches. The Center uniquely supports simultaneous measurement of hundreds to thousands of RNA targets—up to whole-transcriptome profiling—as well as high-plex protein detection ranging from tens to over a thousand markers in a single experiment. This depth of multiplexing allows comprehensive mapping of cellular states, signaling pathways, and cell–cell interactions within complex tumor and tissue microenvironments. By integrating advanced spatial platforms with expert study design, standardized workflows, and end-to-end analytical support, the center enhances rigor and reproducibility and accelerates high-impact basic, translational, and clinical research across St. Jude.
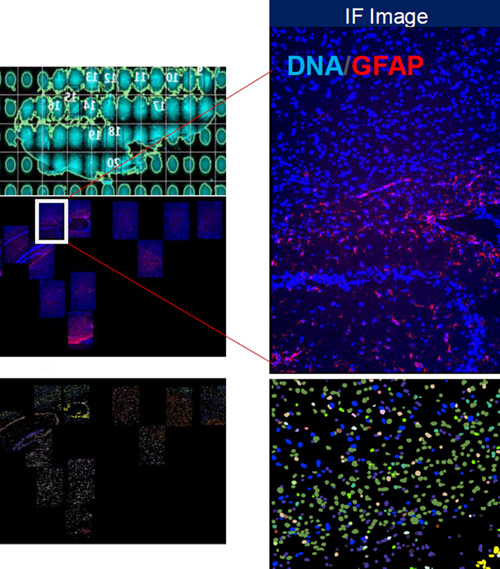
Supervised cell type clustering applied to results generated by RNA/ISH-based CosMx for mouse brain sample.
Equipment/Services
The center provides access to leading technologies for data collection and analysis, including high-level multiplexing and next-generation sequencing, along with advanced computational pipelines, data integration, and multi-omics analyses.
Sequencing-based spatial
Spatially profile the whole transcriptome
Imaging-based spatial
High-resolution imaging with targeted probe design
Multi-Omics
Explore same slidetranscriptomics, proteomics, and more
Data analysis
Bioinformatic analysis available upon request
Data analysis services
- Primary processing and pipeline development
- Automated, high-performance pipelines for image processing, cell segmentation, and molecular quantification across technologies.
- Quality control (QC)
- Filtering and normalization, monitoring metrics such as feature counts, UMI distributions, and tissue integrity to ensure data reliability.
- Cell-type deconvolution and annotation
- Integration of spatial transcriptomics with single-cell RNA-seq (scRNA-seq) references to resolve cell identities and map their precise locations. This includes both reference-based label transfer and de novo annotation of novel cellular states within the tissue architecture.
- Advanced spatial analytics
- Investigation of the "spatial connectome," including cell-cell interaction analysis, neighborhood/niche identification, and spatial trajectory inference.
- Multi-omic integration
- Computational strategies to merge transcriptomic, proteomic, and epigenetic layers, providing a unified view of the tissue microenvironment.
- Interactive visualization and support
- Delivery of data in user-friendly formats and interactive dashboards, coupled with expert consultation to help investigators interpret complex spatial patterns.
Available spatial platforms
- Illumina Spatial Sequencing
- CSO’s newest sequencing-based assay offers whole transcriptome profiling at high resolution. Now supporting fresh frozen tissue that can fit within the 50 x 15 mm capture area, with support for FFPE and fixed frozen tissue in development.
- GeoMx Digital Spatial Profiler (Bruker Spatial Biology)
- Enables highly multiplexed, spatially resolved profiling of RNA (whole transcriptome) and protein (1200 targets) from tissue sections using region-of-interest (ROI) selection. Supports flexible experimental design for discovery and targeted studies and includes custom antibody testing services.
- Visium HD (10x Genomics)
- Provides whole-transcriptome sequencing with high spatial resolution on FFPE tissue sections. Ideal for unbiased transcriptomic mapping across tissue architecture and supports custom probe spike-in services to enhance assay flexibility.
- CosMx Spatial Molecular Imager (Bruker Spatial Biology)
- Delivers single-cell and subcellular resolution for spatial RNA (up to ~6,000 genes) and high-plex protein detection, enabling true same-cell multiomic analysis. Includes custom RNA probe and antibody design services.
- Xenium In Situ (10x Genomics)
- An imaging-based platform for high-plex RNA detection (up to ~5,000 gene targets) with subcellular resolution, with optional integrated protein analysis. Supports custom probe design for targeted and pathway-focused studies.
- PhenoCycler Fusion (Akoya Biosciences)
- Designed for highly multiplexed spatial protein imaging, enabling deep phenotyping of tissue architecture and cellular interactions. Includes pre-designed panel of up to 60 antibodies and custom antibody conjugation services for up to 5 antibodies on one slide.
- Custom Panel and Antibody Design
- Custom probe and/or antibody panels can be designed for all platforms listed. Pricing for custom panels varies by company. Custom design panels require at least a 6-week lead time and additional tissue for optimization.
Selected publications
About the Director
-
View Details
Jasmine Plummer, PhD
Associate Member, St. Jude Faculty
Director, Center for Spatial OMICs
Jasmine Plummer, PhD, is the founding Director of the Center for Spatial Omics and an Associate Member in the Department of Developmental Neurobiology. Her postdoctoral research focused on genetic risk in neurodevelopmental disorders. Dr. Plummer specializes in using multi-omics to study genetic risk in cancers and neurodevelopmental disorders and is known for developing spatial omics platforms.
Jasmine Plummer, PhD
Associate Member, St. Jude Faculty
Director, Center for Spatial OMICs
Affiliations
Research Interests
- Single cell systems biology
- Spatial genomics and proteomics
- Disease pathogenesis from cells of origin
- Genomic technology development
Short Biography
Jasmine Plummer, PhD, is the founding Director of the Center for Spatial Omics and an Associate Member in the Department of Developmental Neurobiology. Her postdoctoral research focused on genetic risk in neurodevelopmental disorders. Dr. Plummer specializes in using multi-omics to study genetic risk in cancers and neurodevelopmental disorders and is known for developing spatial omics platforms.
Contact Information
Jasmine Plummer, PhD
Developmental Neurobiology
MS 322
St. Jude Children's Research Hospital
262 Danny Thomas Place
Memphis, TN 38105-3678
Meet the team
-
View Details
Arjumand Wani, PhD
Scientist
-
View Details
Hannah Chasteen, MS
Lead Researcher
-
View Details
Felipe Segato Dezem
Sr. Bioinformatics Analyst
-
View Details
Maycon Marção, MS
Bioinformatics Analyst





