St. Jude Family of Websites
Explore our cutting edge research, world-class patient care, career opportunities and more.
St. Jude Children's Research Hospital Home

- Fundraising
St. Jude Family of Websites
Explore our cutting edge research, world-class patient care, career opportunities and more.
St. Jude Children's Research Hospital Home

- Fundraising
Lead Discovery Informatics Center
A world-class collaborative research center with expertise in chemical informatics, molecular simulation, computer-aided molecular design, and chemically aware machine learning.
About the Center

The Department of Chemical Biology and Therapeutics (CBT) founded the Lead Discovery Informatics (LDI) in 2006 and has developed it into a world-class collaborative research center with expertise in chemical informatics, molecular simulation, computer-aided molecular design, and chemically aware machine learning. We help build and maintain the chemocentric informatics infrastructure of St. Jude.
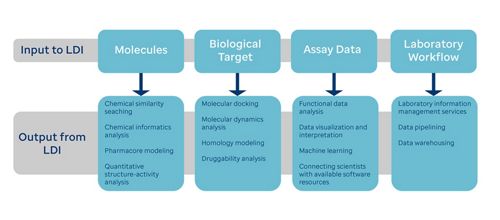
Our mission is to (a) develop and apply algorithms to model molecular interactions and drug efficacy; (b) design and implement tools for visualizing and analyzing molecular properties and relationships; (c) build robust workflows and data pipelines for lab automation; and (d) deploy, customize, and maintain laboratory information management systems.
LDI contributes to projects both scientifically and technically. Scientific contributions include applying computational chemistry methods to analyze ligand binding, writing a program to analyze and visualize data in novel ways, and using chemical informatics approaches to study structure-activity relationships (SAR) and identifty similarities between molecules. Technical contributions include assisting with the installation of chemicentric software, upgrading enterprise applications to maintain compliance with institutional standards, conducting software training sessions, and debugging custom software applications.
Center Staff
-
View Details
Matthew N. Anyanwu, PhD
Sr Scientific Computing Engineer
-
View Details
Sourav Das, PhD
Sr Cheminformatics Research Scientist
-
View Details
Jason M. Ochoada, MS
Cheminformatics Research Scientist
-
View Details
Anang A. Shelat, PhD
Associate Member, St. Jude Faculty
Director, Lead Discovery Informatics
-
View Details
Nathaniel R. Twarog, PhD
Sr Cheminformatics Research Scientist
Contact us
Anang A. Shelat, PhD
Chemical Biology & Therapeutics
MS 1000, Room E9064
St. Jude Children Research Hospital
Follow Us




