St. Jude Family of Websites
Explore our cutting edge research, world-class patient care, career opportunities and more.
St. Jude Children's Research Hospital Home

- Fundraising
St. Jude Family of Websites
Explore our cutting edge research, world-class patient care, career opportunities and more.
St. Jude Children's Research Hospital Home

- Fundraising
McKinnon Lab
Investigating how maintenance of genome stability prevents nervous system disease
About the McKinnon Lab
Genome instability often plays a primary, or confounding, role in certain neurological disorders. Our research program is focused on understanding types of DNA damage and genome maintenance pathways related to chromatin structure. We are also interested in deciphering specific cell type susceptibility as it relates to genome stability and neurological disease pathogenesis. Our work is poised to inform disease mechanism and potential points of intervention.
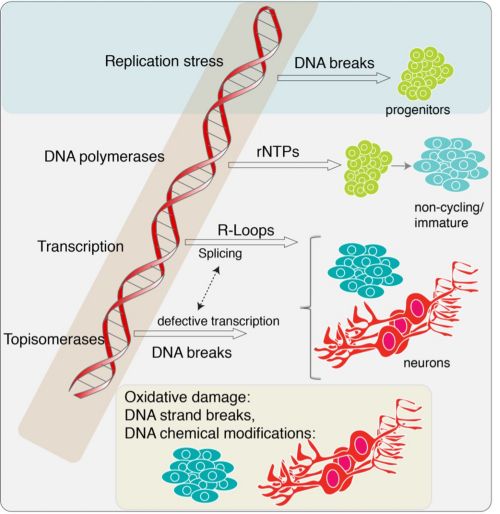
Our research summary
Several inherited human syndromes result from mutations in factors that contribute to genome stability and often result in catastrophic neurodegenerative diseases. Genome instability has also been linked to broader neurological health issues. Our laboratory studies how maintainance of genome integrity prevents diseases of the nervous system. An in-depth knowledge of the processes underlying genome stability is critical to understanding the development and appropriate function of the nervous system. We are particularly interested in understanding 1) which endogenous DNA lesions contribute to pathogenicity, 2) why DNA damage selectively induces tissue-specific pathogenicity, and 3) in contrast to neurogenesis, what DNA damage responses are required by differentiated neural cells in the mature nervous system.

A primary focus in our laboratory involves understanding cell type specificity within the nervous system, particularly the cerebellum, and creating an understanding of how different lesions in particular cell types lead to distinct diseases. We are leveraging genomic approaches including RNA-seq, exome sequencing, ChIP-seq, and ATAC-seq to understand how the chromatin structure of Purkinje cells, oligodendrocytes, motor neurons, and other cell types influences genome stability. In addition, our laboratory is exploring the impact of DNA damge in glia, a critical yet often understudied cell population and their role in neurodegeneration. We ultimately aim to understand the impact of genome damage in specific neural cell types and how this contributes to neurological disease pathogenesis.
Emerging pathology can be difficult to track in disease models. As such, we are using single cell analysis of neurological disease models to provide a high-resolution picture of the impact of various DNA damage events in the early stages of disease development. This approach will capture information often lost by bulk RNA-seq and similar ‘averaged’ approaches and provide a spatiotemporal perspective on susceptibility and potential vulnerabilities in individual neural cell types.
Selected Publications
Contact us
Peter J. McKinnon, PhD
Genomic and Translational Neuroscience
MS 331
St. Jude Children's Research Hospital

Memphis, TN, 38105-3678 USA GET DIRECTIONS